Summary Network¶
A Summary Network is a new network where each node represents a collapsed cluster in the original network, and each edge represents a meta-edge between clusters. The resulting network is very similar to the results you get from collapsing the clusters.
| Before | After |
|---|---|
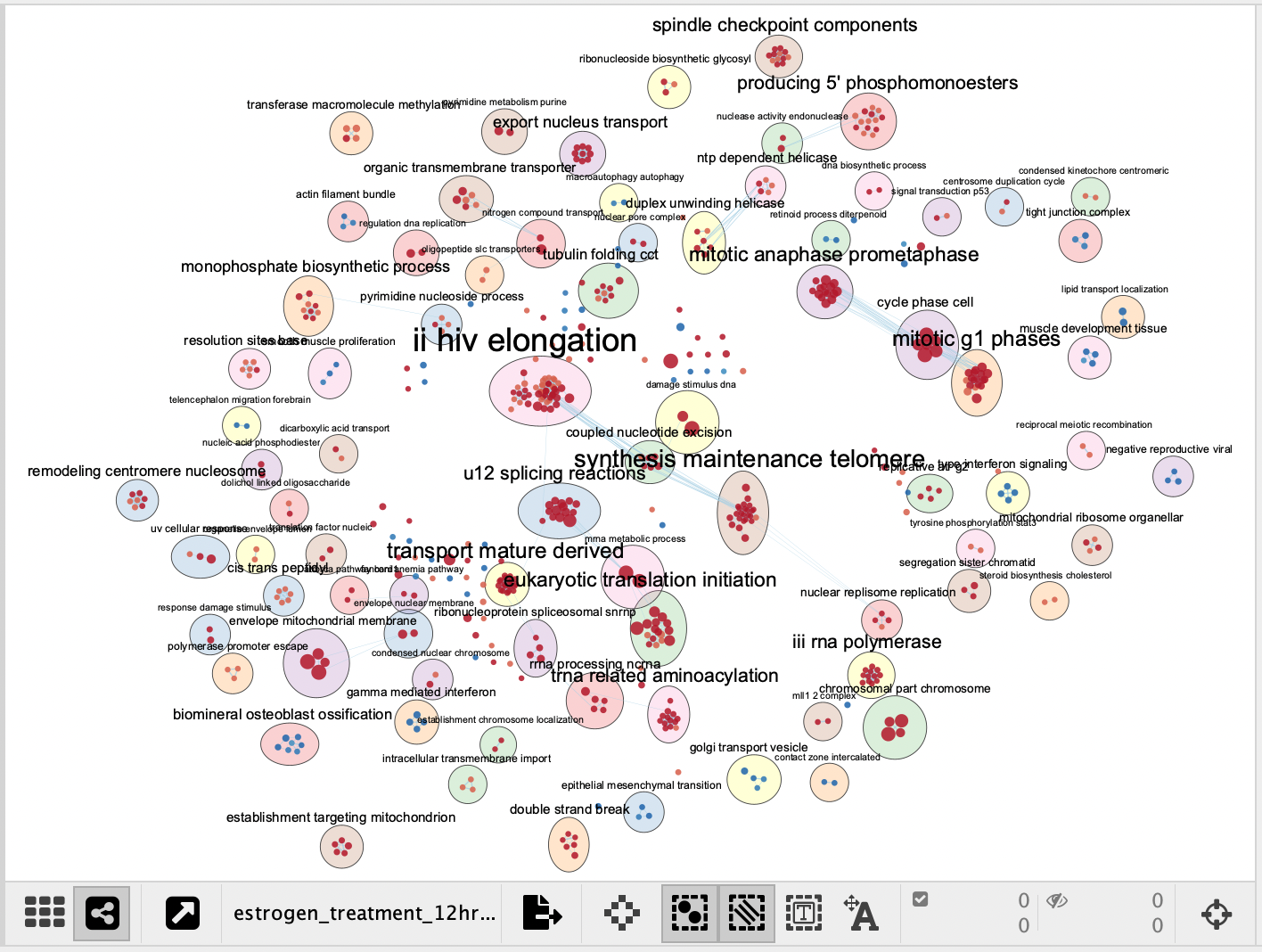 |
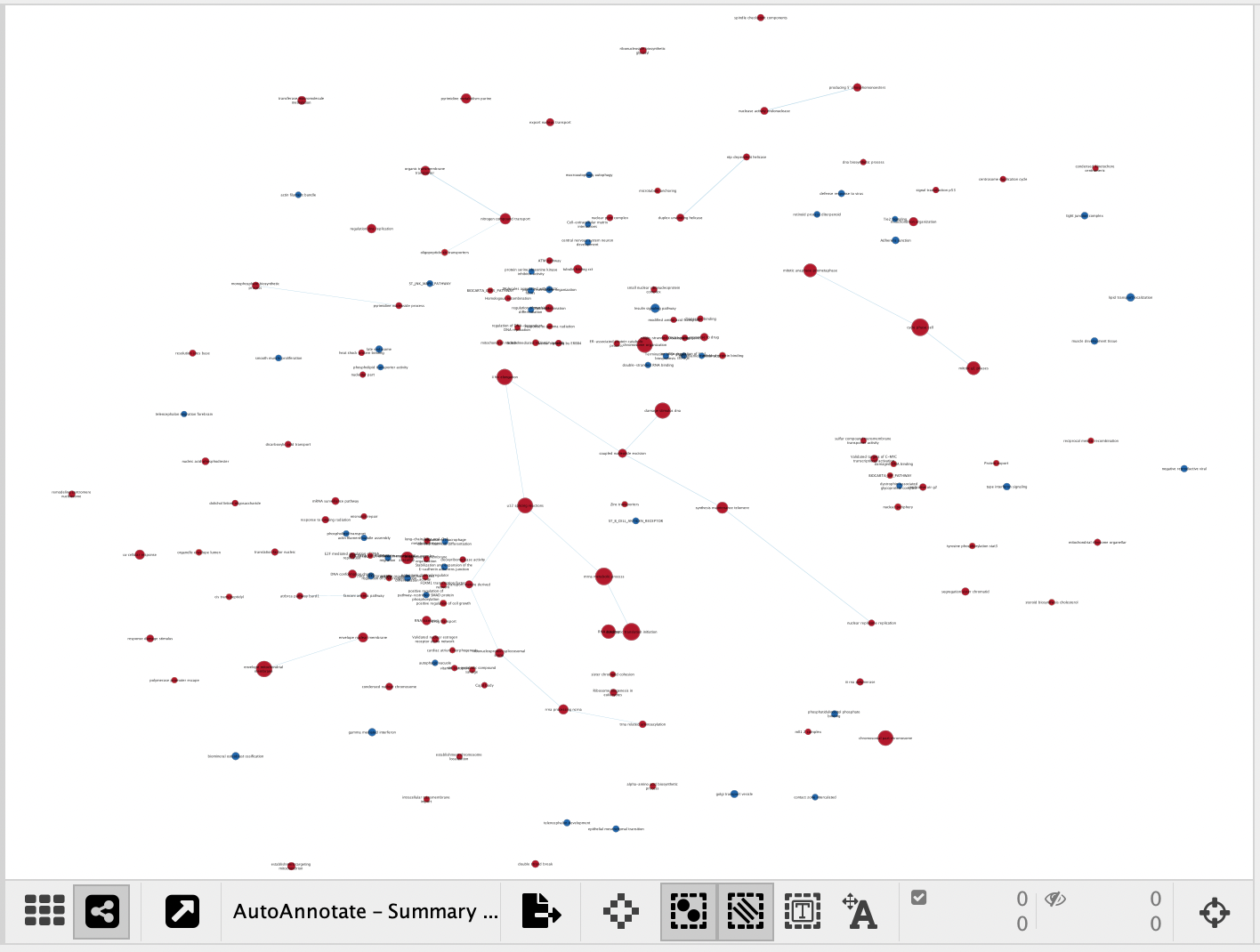 |
A summary network can be created from the Annotation Set Menu or the Cluster Table Context Menu.
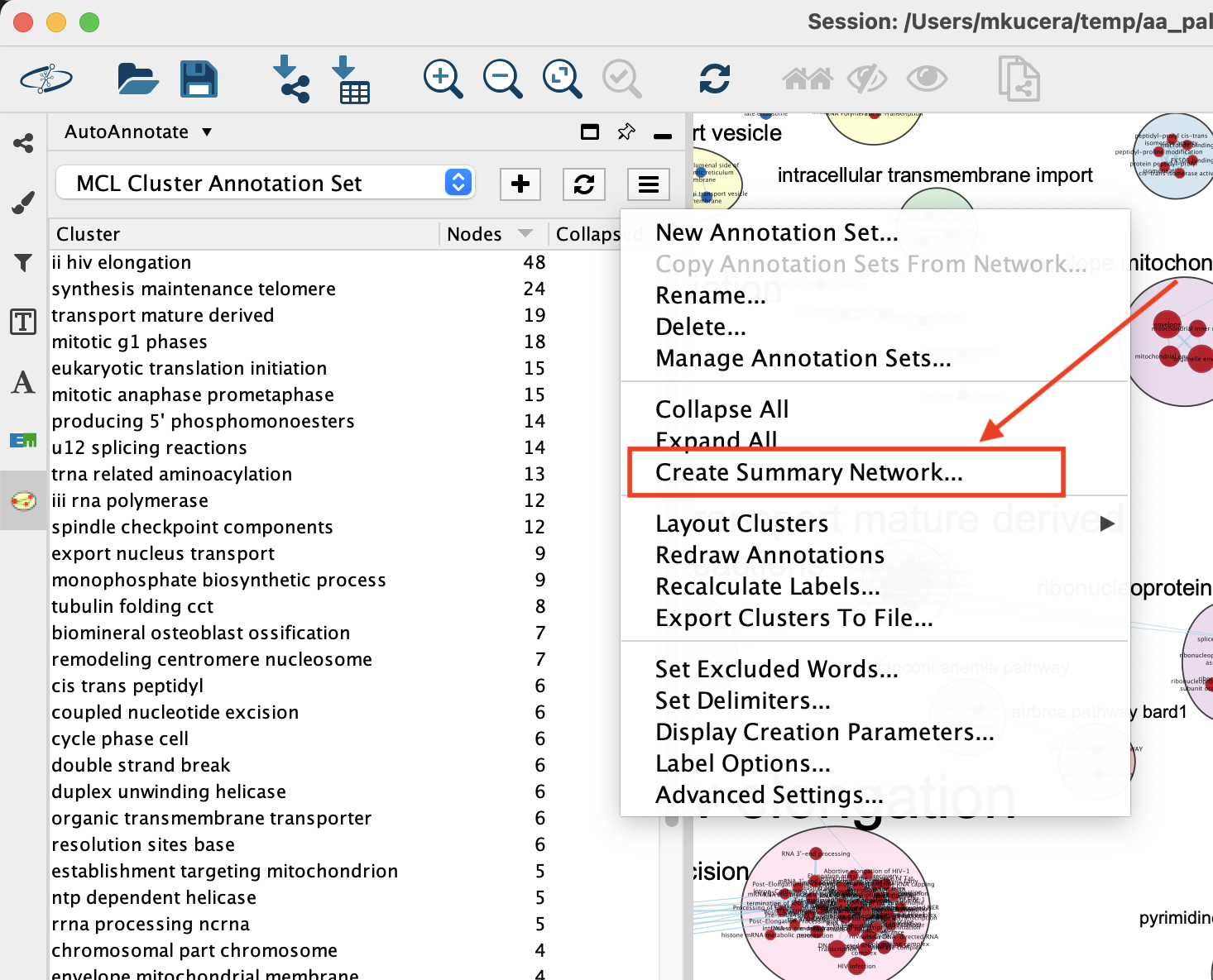
Note
The terms “attribtue” and “column” are used interchangeably.
Attribute Aggregation¶
Each cluster in the source network will be compressed into a single node in the Summary network. If there are multiple edges between two clusters, they will be compressed into a single meta-edge.
The nodes in a cluster may have different values for each data attribute (i.e. column). When the nodes in a cluster are compressed into a single node, the values of the attriutes of the original nodes need to be combined into a single value. There are many options for how this can be done. The same is also true for the edges that are compressed into a meta-edge.
When creating a summary network a dialog is shown that allows you to choose the aggregation strategy for each attribute. Sensible defaults are chosen, so you may safely click OK. However if you would like to customize how attribtues are aggregated use the combo boxes next to each attribute name.
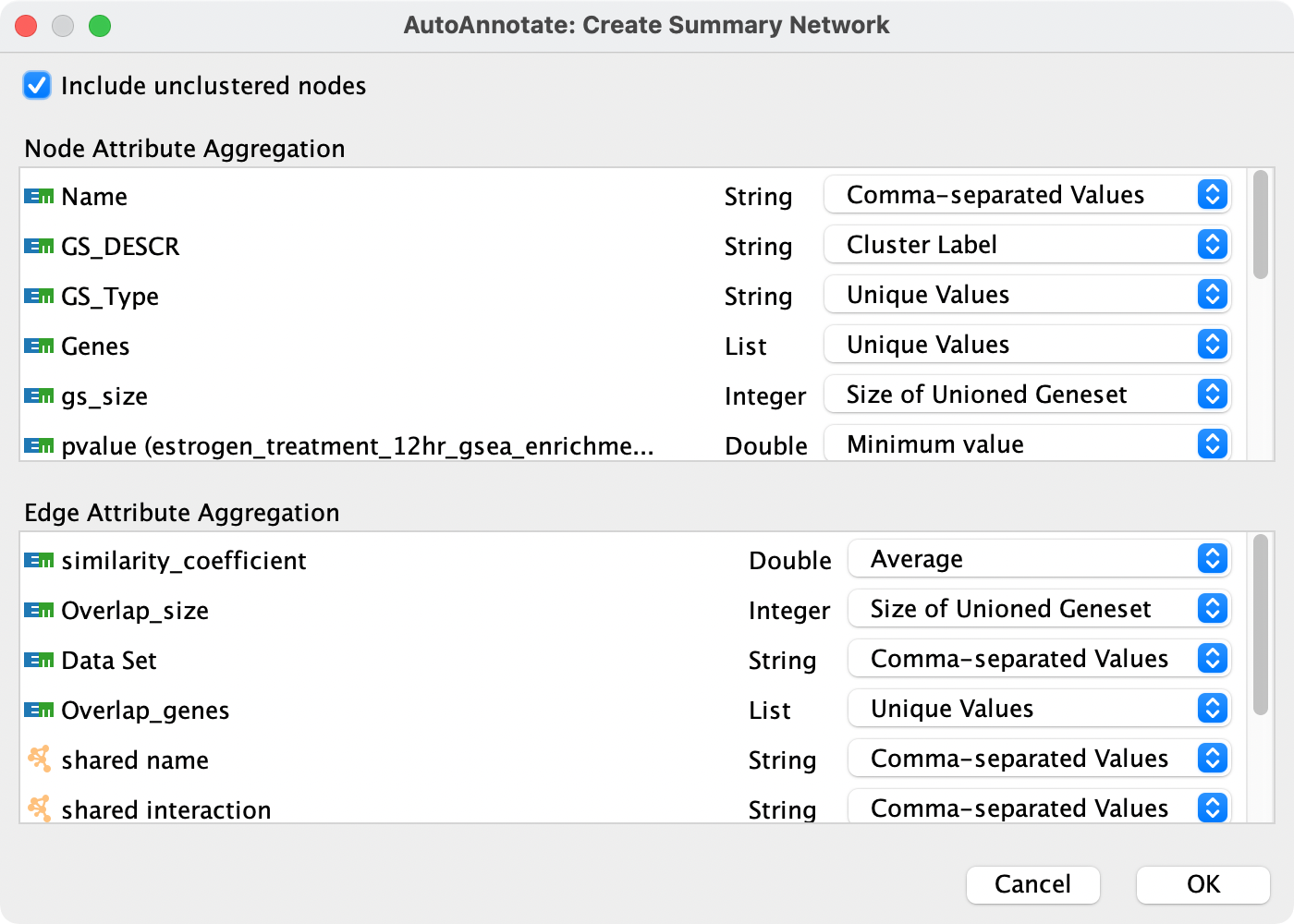
Types of Aggregators¶
None
The resulting value will always be blank.
Comma Separated Values, Tab Separated Values
The values will be concatenated together using commas or tabs as delimiters.
Most Common Value
The most common value will be chosen. Ties are broken arbitrarily.
Unique Values
Takes all the values for an attribute, removes the duplicates, then concatenates the results (using commas as delimiters). For example, a set of nodes with values"a", "a", "a", "b", "b", "c"would become a string with the value"a,b,c".
Cluster Label
Ignores the values of the attribute and uses the cluster label value. Note: If you want a new attribute (column) that contains the cluster label then you must create a new column in the source network before creating the summary network.
Most Significant Gene Set (EnrichmentMap only)
The resulting value is the name of the gene set that has the most significant FDR q-value. If there are ties then the result is the concatenation of the gene set names.
Average, Minimum, Maximum, Median, Sum (Numeric Attributes)
Combines numeric values using the corresponding mathematical operator.
Size of Unioned Geneset (EnrichmentMap only)
Nodes in EnrichmentMap networks represent gene sets. The resulting value is the size of the union of all the gene sets in the cluster.
Greatest Magnitude (EnrichmentMap only)
The result is the value that has the greatest absolute value (i.e. magnitude). For example the set-1, 0, 1, 2would result in2, and the set-2, -1, 0, 1would result in-2.
Logical AND, Logical OR
Used to combine boolean values.
Concatenate (Lists only)
Concatenates the values of list columns. For example[1,2,3]and[4,5,6]would result in[1,2,3,4,5,6].